Overview: GeneMarker HTS - mtDNA Analysis Software for NGS Data
Product informationLicense Options Trial Request Technical Support
mtDNA Analysis Software for (High Throughput Sequencing) NGS Data
GeneMarker®HTS software provides a streamlined workflow for forensic and medical research mitochondrial DNA data analysis from massively parallel sequencing (MPS) systems such as the Illumina® and Ion Torrent® platforms; in an easy to use Windows® operating system. Developed in collaboration with leading laboratories, GeneMarkerHTS software provides rapidanalysis of multiple samples using consensus alignment or a unique motif alignment technology that automates the recommendations of DNA Commission of the International Society for Forensic Genetics: Revised and extended guidelines for mitochondrial DNA typing. Using forensic motif alignment fulfills the maximum parsimony approach of forensic alignment, and provides recognition and proper assignment of motifs and INDELs consistent with phylogenetic and forensic considerations.
 Analysis results include:
Analysis results include:
- Consensus sequence, Variants, SNPs, Indels
- Depth of coverage graphics
- Consensus sequence aligned to reference (IUPAC nomenclature)
- Whole mtDNA genome, spanning the origin
- Specified areas of interest, such as control region, HV1, HV2
- Read pile-up (with depth and direction indicators)
- Compare multiple samples in single view
- Synchronized view, scroll and zoom of multiple samples
- Comparison viewer, table with sample-to-sample and variant composition
Software can export a variety of reports:
- Consensus sequence
- Variant reports – SNPs, insertions and deletions
- Haplotype, heteroplasmy
- Report compatible for import into EMPOP (EDNAP mtDNA Population Database)
The software includes:
- Customizable viewing and reporting to protect privacy of potential health information (PHI)
- Audit trail capability
- User management
- Comparison Capabilities
The initial release of GeneMarkerHTS provides rapid analysis, comparisons and reporting for forensic mtDNA Next Generation data from Illumina and Ion Torrent platforms. Subsequentreleases will include analysis of targeted amplicons from nuclear DNA, such as STRs, identity SNPs and mixture analysis.
Global and zoom example of a whole mtDNA genome alignment:
The Global View shows the depth of coverage with forward read coverage in blue and reverse read coverage in red.
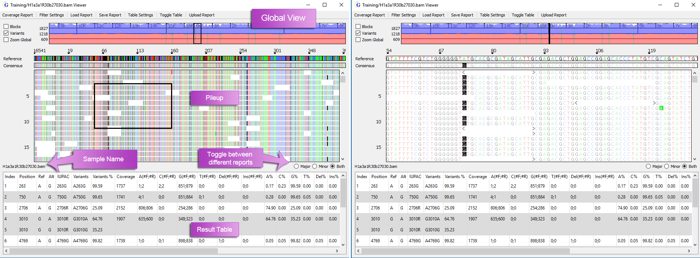
Rapid Analysis and Synchronized Viewing of Multiple Samples
GeneMarkerHTS software provides results in minutes; for example, 30 MiSeq whole mtDNA chromosome data files with 10,000 average depth of coverage were aligned in 90 minutes (3 minutes per sample). In a more extreme example, 200 whole mtDNA chromosome data files with 10,000 average depth of coverage aligned in 16 hours.
Information from Whole mtDNA genome is rapidly analyzed, or choose to analyze specific regions, such as the control region or HV1 & HV2 analysis in minutes with these automatic features:
- Analysis across the origin
- Forensic or standard nomenclature
- Maintain health privacy with an administrative option that blocks display of disease associated positions
- Three alignment algorithms that best suit the analysis needs:
- Initial Alignment to a reference genome (Revised Cambridge Reference Sequence is preloaded into GeneMarkerHTS)
- Adjust alignment with motifs to ensure the alignment is consistent with forensic variant reporting
- Adjust alignment using the consensus; improving alignment around indels
- Customize viewing options
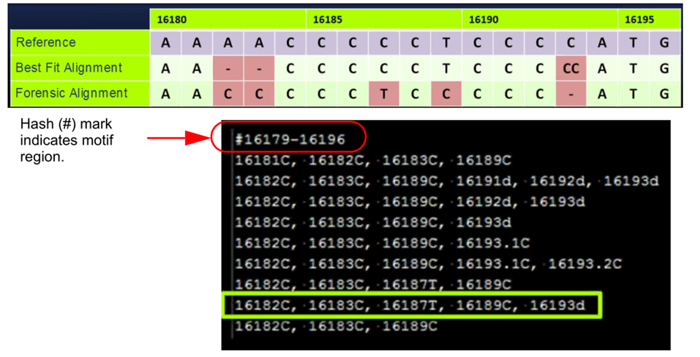
Motif alignment reduces manual edits for forensic alignment
GeneMarkerHTS software has an extensive, preloaded forensic motif file as well as a motif editor to assist labs in adding new motifs.
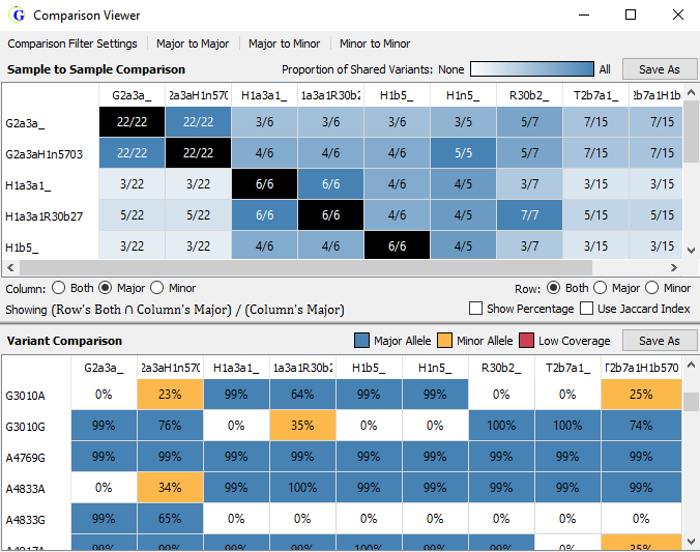
Additional Tools:
Comparison Viewer is a viewing tool to compare analysis results of multiple samples. Use the comparison viewer for:
- Sample to sample comparison
- As well as a variant comparison of all the samples in the project at the same time
Audit Trail, User Management and Database functions:
The lab administrator establishes user ID with access rights. The audit trail provides a history of permissions, project and user activities.
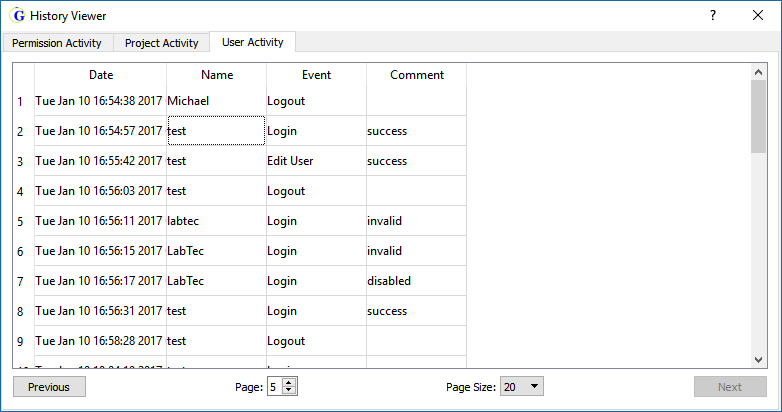
The database allows users to upload/download projects with the initial analysis parameters and upload/download changes to analysis parameters of subsequent analysts. The database provides a record of all analysis parameters and activities on a data set
Read the International Journal of Legal Medicine Article
GeneMarker HTS Introductory Webinar
Technical Support
Trial, Quote & Webinar Request Form
The supporting documents available for this product can be downloaded below.