Find your product
Advanced searchLogin
Webshop under construction
Due to technical maintenance the webshop is closed until January 3, 4. We wish you a successful 2024!
Welcome
Sign outPersonal information
99.8% Accuracy with Ion PGM™ PE reads and NextGENe software’s Floton™ Merge Tool
The Ion PGM system now provides a paired end sequencing option in which DNA fragments can be sequenced from both directions. Paired end sequences can be merged into a single, high-quality sequence using NextGENe’s “Overlap Merger” tool. This process greatly improves accuracy, especially at the 3’ end, reducing the need for trimming. This approach makes it possible to improve the accuracy of the Ion Torrent platform to more than 99.8% when the “hide unmatched ends” option is used. This is especially useful for AmpliSeq™ projects.
Bacterial Dataset Summary
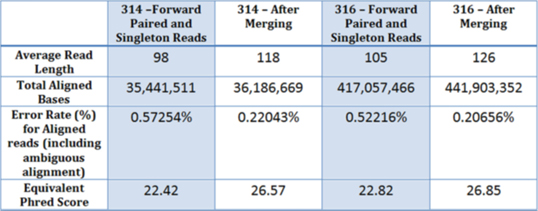
Read Length and Aligned Read Accuracy Improvements for two bacterial re-sequencing projects. The adapter sequence (GCTGAGGA) was trimmed for the non-merged data.
AmpliSeq™ Dataset Summary
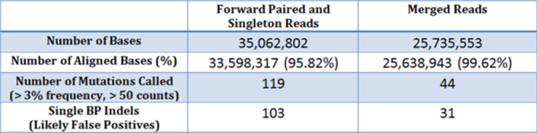
Alignment and mutation calling comparison of an AmpliSeq dataset. For the purposes of comparison, mutations called in the Amplicon regions were only filtered with an allele count and mutant percentage filter.
Paired End Sequencing data from the Ion PGM provides improved accuracy by sequencing the same fragment from each direction. In addition to greater accuracy there are several other benefits including: Adapter sequences are removed, both ends of the merged sequences have high quality basecalls , reads are longer on average because quality trimming is not needed, larger fraction of reads are able to align and a lower false positive rate.
To learn more or to obtain a 30-day NextGENe trial email: [email protected]