EcoP15I
Product information| Code | Name | Size | Quantity | Price | |
|---|---|---|---|---|---|
R0646S |
EcoP15I, recombinant |
500 units ( 10000 units/ml ) | - | Unavailable in your region |
EcoP15I
We are excited to announce that all reaction buffers are now BSA-free. NEB began switching our BSA-containing reaction buffers in April 2021 to buffers containing Recombinant Albumin (rAlbumin) for restriction enzymes and some DNA modifying enzymes. Find more details at www.neb.com/BSA-free.

Product Introduction
- This product is supplied with one vial of 10X ATP solution (10mM concentration, NEB #B6101S, not sold separately)
- Time-Saver™ qualified for digestion in 5-15 minutes
-
Requires two or more sites for cleavage. Please review the enzyme list and recommendations in the following table as well as the background information article.
-
Restriction Enzyme Cut Site: CAGCAG(25/27)
| Catalog # | Size | Concentration |
|---|---|---|
| R0646S | 500 units | 10000 units/ml |
| R0646L | 2500 units | 10000 units/ml |
Featured Videos
View Video Library-

Reduce Star Activity with High-Fidelity Restriction Enzymes
-
TIME-SAVER™ Protocol for Restriction Enzyme Digests
-

NEB® TV Ep. 15 – Applications of Restriction Enzymes
-

Restriction Enzyme Digest Protocol: Cutting Close to DNA End
-

Restriction Enzyme Digestion Problem: DNA Smear on Agarose Gel
-
Why is My Restriction Enzyme Not Cutting DNA?
-

Restriction Enzyme Digest Problem: Too Many DNA Bands
-
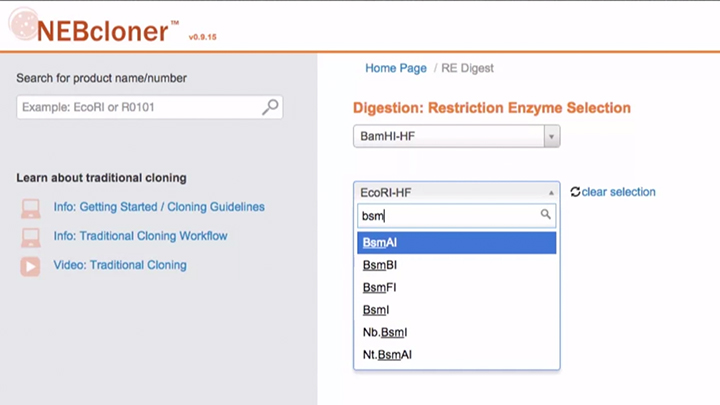
Double Digestion with NEBcloner
- Product Information
- Protocols, Manuals & Usage
- Tools & Resources
- FAQs & Troubleshooting
- Citations & Technical Literature
- Quality, Safety & Legal
- Other Products You May Be Interested In
Product Information
Description
Product Source
An E. coli strain that carries the cloned EcoP15 l res-mod genes from plasmid pSHl180 (D.N. Rao)- This product is related to the following categories:
- Restriction Endonucleases C G,
- Time-Saver Qualified Restriction Enzymes,
- This product can be used in the following applications:
- Fast Cloning: Accelerate your cloning workflows with reagents from NEB,
- Restriction Enzyme Digestion
Reagents Supplied
Reagents Supplied
The following reagents are supplied with this product:
| NEB # | Component Name | Component # | Stored at (°C) | Amount | Concentration | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ||||||||||||||||||||||||||||||
Properties & Usage
Unit Definition
One unit is defined as the amount enzyme required to digest 1 µg of pUC19 DNA in 1 hour at 37°C in a total reaction volume of 50 µl.Reaction Conditions
1X NEBuffer™ r3.1
Supplement with 1X ATP
Incubate at 37°C
1X NEBuffer™ r3.1
100 mM NaCl
50 mM Tris-HCl
10 mM MgCl2
100 µg/ml Recombinant Albumin
(pH 7.9 @ 25°C)
Activity in NEBuffers
NEBuffer™ r1.1: 75%NEBuffer™ r2.1: 100%
NEBuffer™ r3.1: 100%
rCutSmart™ Buffer: 100%
Diluent Compatibility
Storage Buffer
10 mM Tris-HCl
100 mM NaCl
1 mM DTT
0.1 mM EDTA
200 µg/ml BSA
50% Glycerol
pH 7.4 @ 25°C
Heat Inactivation
65°C for 20 minutesMethylation Sensitivity
dam methylation: Not Sensitive
dcm methylation: Not Sensitive
CpG Methylation: Not Sensitive
Product Notes
- ATP is required for DNA cleavage. Efficient cleavage requires the presence of two inversely oriented recognition sites. A head to head orientation is preferred. The "head" of the sequence is defined as the dG at the 3´ end of the CAGCAG strand (top strand). Cleavage efficiency is also affected by the distance between the two sites.
- EcoP15I requires two copies of its recognition sequence for cleavage to occur, with limited cleavage on plasmids with a single site.
- Based on the stability of the enzyme in the reaction, incubations longer than 1 hr will not result in improved digestion, unless additional enzyme is added. Please refer to Restriction endonuclease survival in a reaction for more information regarding this topic.
- Requires two or more sites for cleavage. Please review the enzyme list and recommendations in the following table as well as the background information article.
- Not sensitive to CpG, dcm, or dam methylation.
References
- Elisabeth Möncke-Buchner, Maja Rothenberg, Stefanie Reich, Katja Wagenführ, Hideo Matsumura, Ryohei Terauchi, Detlev H. Krüger, and Monika Reuter (2009). Functional Characterization and Modulation of the DNA Cleavage Efficiency of Type III Restriction Endonuclease EcoP15I in Its Interaction with Two Sites in the DNA Target. J. Mol. Biol. 387, 1309–1319.
Protocols, Manuals & Usage
Protocols
Usage & Guidelines
- Activity at 37°C for Restriction Enzymes with Alternate Incubation Temperatures
- Activity of Restriction Enzymes in PCR Buffers
- Cleavage Close to the End of DNA Fragments
- Digestion of Agarose-Embedded DNA: Info for Specific Enzymes
- Double Digests
- Heat Inactivation
- NEBuffer Activity/Performance Chart with Restriction Enzymes
- Optimizing Restriction Endonuclease Reactions
- Restriction Endonucleases - Survival in a Reaction
- Restriction Enzyme Diluent Buffer Compatibility
- Restriction Enzyme Tips
- Restriction enzymes requiring multi-sites for efficient cleavage
- Single Letter Codes
- Star Activity
- Traditional Cloning Quick Guide
Tools & Resources
Selection Charts
- Alphabetized List of Recognition Sequences
- Compatible Cohesive Ends and Generation of New Restriction Sites
- Dam-Dcm and CpG Methylation
- Enzymes with Nonpalindromic Sequences
- Frequencies of Restriction Sites
- Isoelectric Points (pI) for Restriction Enzymes
- Isoschizomers
- NEB Diluent and Buffer Table
- Why Choose Recombinant Enzymes?
Web Tools
FAQs & Troubleshooting
FAQs
Troubleshooting
Citations & Technical Literature
Citations
Additional Citations
- Bhattacharyya, S., Yu, Y., Suzuki, M., Campbell, N., Mazdo, J., Vasanthakumar, A., et al. (2013) Genome-wide hydroxymethylation tested using the HELP-GT assay shows redistribution in cancer Nucleic Acids Res; DOI: doi:10.1093/nar/gkt601
Quality, Safety & Legal
Quality Assurance Statement
Quality Control tests are performed on each new lot of NEB product to meet the specifications designated for it. Specifications and individual lot data from the tests that are performed for this particular product can be found and downloaded on the Product Specification Sheet, Certificate of Analysis, data card or product manual. Further information regarding NEB product quality can be found here.Specifications
The Specification sheet is a document that includes the storage temperature, shelf life and the specifications designated for the product. The following file naming structure is used to name these document files: [Product Number]_[Size]_[Version]Certificate Of Analysis
The Certificate of Analysis (COA) is a signed document that includes the storage temperature, expiration date and quality controls for an individual lot. The following file naming structure is used to name these document files: [Product Number]_[Size]_[Version]_[Lot Number]- R0646S_L_v1_0151403
- R0646S_L_v1_0151301
- R0646S_L_v1_0151306
- R0646S_L_v1_0151308
- R0646S_L_v1_0151312
- R0646S_L_v1_0151405
- R0646S_L_v1_0151407
- R0646S_L_v1_0211410
- R0646S_L_v1_0211412
- R0646S_L_v2_0221507
- R0646S_L_v2_0221509
- R0646S_L_v2_0221511
- R0646S_L_v2_0221604
- R0646S_L_v2_0221607
- R0646S_L_v2_0221611
- R0646S_L_v2_0221701
- R0646S_L_v2_0221710
- R0646S_L_v2_0221803
- R0646L_v2_10007480
- R0646L_v2_10020415
- R0646S_v2_10025266
- R0646L_v2_10039322
- R0646S_v2_10039324
- R0646L_v2_10050218
- R0646S_v2_10050215
- R0646L_v2_10060442
- R0646S_v2_10060441
- R0646S_v2_10070478
- R0646S_v2_10078793
- R0646S_v2_10082953
- R0646S_v2_10090889
- R0646S_v2_10090941
- R0646S_v2_10101558
- R0646S_v2_10132365
- R0646S_v2_10116810
- R0646S_v2_10150220
- R0646S_v2_10182637
- R0646S_v2_10203027
- R0646S_v2_10231317
- R0646S_v2_10236868
- R0646S_v2_10251750
- R0646S_v2_10264725
- R0646S_v2_10289779
Safety DataSheets
The following is a list of Safety Data Sheet (SDS) that apply to this product to help you use it safely.EcoP15I
NEBuffer™ r3.1
10X ATP
Legal and Disclaimers
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. The use of this product may require the buyer to obtain additional third-party intellectual property rights for certain applications. For more information, please email [email protected].This product is intended for research purposes only. This product is not intended to be used for therapeutic or diagnostic purposes in humans or animals.
New England Biolabs (NEB) is committed to practicing ethical science – we believe it is our job as researchers to ask the important questions that when answered will help preserve our quality of life and the world that we live in. However, this research should always be done in safe and ethical manner. Learn more.
Other Products You May Be Interested In
The supporting documents available for this product can be downloaded below.