microRNA Marker
Product information| Code | Name | Size | Quantity | Price | |
|---|---|---|---|---|---|
N2102S |
microRNA Marker |
100 gel lanes ( 12 ng/µl ) | - | Unavailable in your region |
microRNA Marker
Product Introduction
- Contains 2 synthetic oligonucleotides: 17, 21 and 25 residues
- Convenient, ready-to-load format
- For use with denaturing polyacrylamide gels and Northern blots
| Catalog # | Size | Concentration |
|---|---|---|
| N2102S | 100 gel lanes (0.5 ml) | 12 ng/µl |
Featured Videos
View Video Library- Product Information
- FAQs & Troubleshooting
- Citations & Technical Literature
- Quality, Safety & Legal
- Other Products You May Be Interested In
Product Information
Description
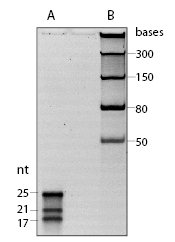
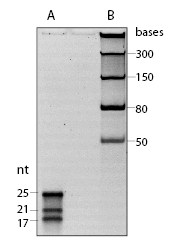
The microRNA Marker is a set of three synthetic single-stranded RNA oligonucleotides 17, 21 and 25 residues long that have free 5´ ends (i.e., no 5´ phosphate groups). These oligonucleotides can be used as size markers on denaturing polyacrylamide gels and Northern blots. The microRNA Marker is best visualized by staining with SYBR-Gold instead of ethidium bromide (Figure 1).The three marker oligos contain the same core sequence so they can be detected by hybridization with the same probe. The sequences of the microRNA marker band are as follows:
25-mer: 5´AGAGCAGUGGCUGGUUGAGAUUUAA 3´
21-mer: 5´AGCAGUGGCUGGUUGAGAUUU 3´
17-mer: 5´CAGUGGCUGGUUGAGAU 3´
Note: The sequence in bold is common to all three oligos.
A 21-mer DNA oligonucleotide complementary to the marker sequences is included. This oligonucleotide is biotinylated at the 3´ end and has a free 5´ end so it can also be labeled with γ-32P-ATP and T4 Polynucleotide Kinase (NEB# M0201). The sequence of the oligonucleotide probe is as follows:
5´AAATCTCAACCAGCCACTGCT 3´-Biotin
The microRNA Marker is provided in a ready-to-load solution containing 4M urea and 0.04% Orange-G. The microRNA Marker Probe is resuspended in water.
- This product is related to the following categories:
- RNA Markers & Ladders,
- RNA Markers and Ladders,
Reagents Supplied
Reagents Supplied
The following reagents are supplied with this product:
| NEB # | Component Name | Component # | Stored at (°C) | Amount | Concentration | |
|---|---|---|---|---|---|---|
Properties & Usage
Related Products
Companion Products
Product Notes
- The microRNA Marker is provided in a ready-to-load denaturing solution. Denature by heating for 3-5 minutes at 95°C and place on ice. Load 5-10 µl for staining with SYBR Gold in denaturing gels. In Northern blots, less than 1 µl (12 ng) is sufficient for detection by hybridization.
The Orange G loading buffer migrates faster than the smallest band, and migrates approximately as far as the nucleotides. - Oligonucleotide Probe Usage: The provided biotinylated probe DNA oligonucleotide can be labeled with T4 Polynucleotide Kinase (NEB #M0201) and radioactive γ-32P-ATP using the following protocol:
1. Mix the following components in a sterile microfuge tube:
Oligonucleotide Probe -- 1-5 µl
10X T4 Polynucleotide Kinase reaction Buffer -- 2.0 µl
γ-32P-ATP (5 µCi/µl) -- 1-2 µl
T4 Polynuclotide Kinase -- 1 µl
Sterile dH20 -- X µl
Total volume -- 20 µl
2. Incubate for 30 minutes at 37°C.
3. Purify labeled probe using a G-25 spin column.
References
- Sambrook, J. and Russel, D.W. (2001). Molecular Cloning: A Laboratory Manual (3rd Ed.). 7.1-7.56.
FAQs & Troubleshooting
FAQs
- Can I use the microRNA Marker on a non-denaturing gel?
- Should I denature the microRNA Marker before loading on my gel?
- What is the difference between the siRNA marker and the microRNA marker (discontinued)?
- Can I use EtBr for staining the microRNA Marker after gel electrophoresis?
- Should I load more of the microRNA Marker so I can get better visibility of the bands after staining the gel with EtBr?
Citations & Technical Literature
Citations
Additional Citations
Quality, Safety & Legal
Quality Assurance Statement
Quality Control tests are performed on each new lot of NEB product to meet the specifications designated for it. Specifications and individual lot data from the tests that are performed for this particular product can be found and downloaded on the Product Specification Sheet, Certificate of Analysis, data card or product manual. Further information regarding NEB product quality can be found here.Specifications
The Specification sheet is a document that includes the storage temperature, shelf life and the specifications designated for the product. The following file naming structure is used to name these document files: [Product Number]_[Size]_[Version]Certificate Of Analysis
The Certificate of Analysis (COA) is a signed document that includes the storage temperature, expiration date and quality controls for an individual lot. The following file naming structure is used to name these document files: [Product Number]_[Size]_[Version]_[Lot Number]Safety DataSheets
The following is a list of Safety Data Sheet (SDS) that apply to this product to help you use it safely.microRNA Marker
Legal and Disclaimers
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. The use of this product may require the buyer to obtain additional third-party intellectual property rights for certain applications. For more information, please email [email protected].This product is intended for research purposes only. This product is not intended to be used for therapeutic or diagnostic purposes in humans or animals.
New England Biolabs (NEB) is committed to practicing ethical science – we believe it is our job as researchers to ask the important questions that when answered will help preserve our quality of life and the world that we live in. However, this research should always be done in safe and ethical manner. Learn more.
Other Products You May Be Interested In
No supporting documents available
This product has no supporting documents available for download. If you feel like supporting documents should be available for this product, please contact us.