Exonuclease V (RecBCD)
Product information| Code | Name | Size | Quantity | Price | |
|---|---|---|---|---|---|
M0345S |
Exonuclease V (RecBCD) |
1.000 units ( 10000 units/ml ) | - | Unavailable in your region | |
M0345L |
Exonuclease V (RecBCD) |
5.000 units ( 10000 units/ml ) | - | Unavailable in your region |
Exonuclease V (RecBCD)
Product Introduction
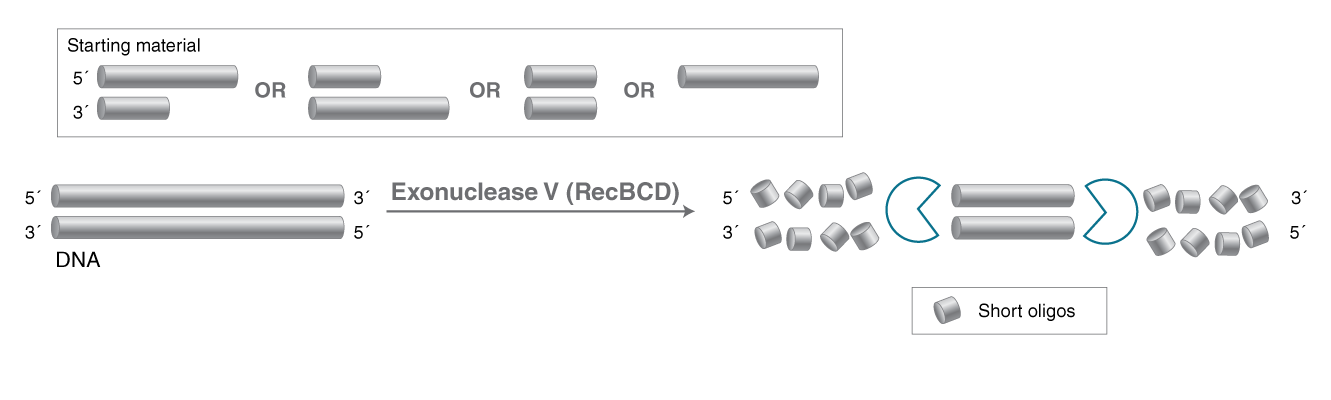
- DNA specific exonuclease that also acts as an endonuclease on single-strand DNA
- Initiates at both 5' and 3' termini of linear double-stranded DNA
- Cleaves linear double-stranded DNA in both the 3' to 5' and 5' to 3' directions
- Requires ATP in the reaction
- Need help finding the right exonuclease for your experiments? Try Exo Selector
Exonuclease V is ideal for:
- Degradation of contaminating linear DNA in plasmid samples
- Removal of residual gDNA after purification of low copy plasmid
| Catalog # | Size | Concentration |
|---|---|---|
| M0345S | 1000 units | 10000 units/ml |
| M0345L | 5000 units | 10000 units/ml |
- Product Information
- Protocols, Manuals & Usage
- Tools & Resources
- FAQs & Troubleshooting
- Citations & Technical Literature
- Quality, Safety & Legal
- Other Products You May Be Interested In
Product Information
Description
Exonuclease V, a RecBCD complex from E. coli has several different enzyme activities, including an ATP-dependent single-stranded DNA endonuclease activity, ss- and ds- DNA exonuclease activity. The hydrolysis in each case is bi-directional (from both the 3´ and 5´ ends) and processive, producing oligonucleotides (1,2,3). All Exonuclease V activities have divalent cation requirements. Mg2+ is required for the exonuclease activity, while Ca2+ inhibits the exonuclease activity and allows double-stranded DNA unwinding (helicase activity) without hydrolysis (4,5).Product Source
An E. coli strain containing plasmids for expressing the three subunits of E. coli Exonuclease V: RecB, RecC and RecD.- This product is related to the following categories:
Reagents Supplied
Reagents Supplied
The following reagents are supplied with this product:
| NEB # | Component Name | Component # | Stored at (°C) | Amount | Concentration | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||
Properties & Usage
Unit Definition
One unit is defined as the amount of enzyme required to produce 1 nmol of acid-soluble deoxyribonucleotide from double-stranded DNA in 30 minutes at 37°C in a total reaction volume of 50 μl.Reaction Conditions
1X NEBuffer™ 4
Supplement with 1 mM ATP
Incubate at 37°C
1X NEBuffer™ 4
50 mM Potassium Acetate
20 mM Tris-acetate
10 mM Magnesium Acetate
1 mM DTT
(pH 7.9 @ 25°C)
Storage Buffer
50 mM Tris-HCl
100 mM NaCl
1 mM DTT
0.1 mM EDTA
50% Glycerol
0.1% Triton® X-100
pH 7.5 @ 25°C
Heat Inactivation
70°C for 30 minutesUnit Assay Conditions
1X NEBuffer 4, 1 mM ATP with 0.15 mM sonicated duplex [3H]-DNA.Application Features
Degradation of linear ssDNA and dsDNA while preserving nicked and supercoiled plasmid DNA.Related Products
Materials Sold Separately
References
- Eichler, D.C. et al. (1977). J.Biol. Chem. 252, 499-503.
- Taylor, A.F. et al. (1985). J. Mol. Biol. 185, 431-443.
- Amundsen, S.K. et al. (1986). Proc. Natl. Acad. Sci. 83, 5558-5562.
- Palas, K.M. et al. (1990). J. Biol. Chem. 265, 3447-3454.
- Dillingham, M.S. et al. (2008). Microbiology and Mol. Biol. Review. 72, 642-671.
Protocols, Manuals & Usage
Protocols
Usage & Guidelines
Application Notes
Tools & Resources
Selection Charts
Web Tools
Research Posters
FAQs & Troubleshooting
FAQs
- What is the best reaction volume for removal of linear dsDNA?
- Does isolated genomic DNA purified by different methods change the cleavage rate of Exonuclease V?
- Should I add additional ATP with Exonuclease V if I am scaling up my reaction?
- Does Exonuclease V (RecBCD) work in other NEBuffers?
- What is the best exonuclease to remove chromosomal DNA during a plasmid prep?
- Which enzyme can distinguish between mitochondrial and genomic DNA and only remove gDNA?
- Can Exonuclease I be used with a double stranded exonuclease to clean up plasmid preparations?
- Does NEB have an equivalent to Lucigen’s Plasmid-Safe?
Citations & Technical Literature
Citations
Additional Citations
Quality, Safety & Legal
Quality Assurance Statement
Quality Control tests are performed on each new lot of NEB product to meet the specifications designated for it. Specifications and individual lot data from the tests that are performed for this particular product can be found and downloaded on the Product Specification Sheet, Certificate of Analysis, data card or product manual. Further information regarding NEB product quality can be found here.Specifications
The Specification sheet is a document that includes the storage temperature, shelf life and the specifications designated for the product. The following file naming structure is used to name these document files: [Product Number]_[Size]_[Version]Certificate Of Analysis
The Certificate of Analysis (COA) is a signed document that includes the storage temperature, expiration date and quality controls for an individual lot. The following file naming structure is used to name these document files: [Product Number]_[Size]_[Version]_[Lot Number]- M0345S_L_v1_0011712
- M0345S_L_v1_0011804
- M0345S_v1_10012582
- M0345L_v1_10018400
- M0345S_v1_10024969
- M0345L_v1_10042754
- M0345S_v1_10045576
- M0345S_v1_10051887
- M0345S_v1_10056008
- M0345L_v1_10051889
- M0345S_v1_10058503
- M0345S_v1_10064165
- M0345L_v1_10067287
- M0345S_v1_10067289
- M0345L_v1_10081095
- M0345S_v1_10088295
- M0345S_v1_10092697
- M0345L_v1_10081097
- M0345L_v1_10105432
- M0345L_v1_10113010
- M0345S_v1_10113011
- M0345L_v1_10137208
- M0345S_v1_10137206
- M0345L_v1_10151813
- M0345S_v1_10160884
- M0345S_v1_10182556
- M0345L_v1_10198648
- M0345S_v1_10203268
- M0345L_v1_10225665
- M0345S_v1_10234127
- M0345L_v1_10238332
- M0345S_v1_10243481
- M0345L_v1_10243480
- M0345L_v1_10248321
- M0345S_v1_10248320
- M0345S_v1_10262432
- M0345L_v1_10267515
- M0345S_v1_10272430
- M0345L_v1_10275133
- M0345S_v1_10280111
- M0345S_v1_10285225
- M0345L_v1_10291484
- M0345S_v1_10291538
Safety DataSheets
The following is a list of Safety Data Sheet (SDS) that apply to this product to help you use it safely.Exonuclease V (RecBCD)
NEBuffer™ 4
Adenosine 5'-Triphosphate (ATP)
Legal and Disclaimers
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. The use of this product may require the buyer to obtain additional third-party intellectual property rights for certain applications. For more information, please email [email protected].This product is intended for research purposes only. This product is not intended to be used for therapeutic or diagnostic purposes in humans or animals.
New England Biolabs (NEB) is committed to practicing ethical science – we believe it is our job as researchers to ask the important questions that when answered will help preserve our quality of life and the world that we live in. However, this research should always be done in safe and ethical manner. Learn more.
Other Products You May Be Interested In
The supporting documents available for this product can be downloaded below.