Please Note
The contents and the protocols of this kit have changed from July 2017! The kit will only contain blue NucleoSpin RNA Columns. The protocols will only show the collection of total RNA in one fraction. There are support protocols available for the fractionation of small and large RNA. Contact us if you have any questions.
|
Principles
NucleoSpin® miRNA is suitable for simultaneous isolation of small RNA (< 200 nt, e.g., miRNA, pre-miRNA, tRNA, 5S RNA), large RNA (> 200 nt, e.g., mRNA, 18S rRNA, 28S rRNA, pri-miRNA), and protein in three separate fractions from a large variety of sample materials.
The precipitated protein can easily be dissolved in Laemmli buffer and used for SDS-PAGE, Western Blot analysis, and protein quantification, for example with the MACHEREY-NAGEL Protein Quantification Assay.
The eluted RNA and miRNA are ready-to-use for all standard downstream applications, for example RT-PCR, Northern Blot, or chip hybridization.
Parallel isolation of small and large RNA
- RNA purification fractionated by size:
Isolation of small RNA only (< 200 b),
Isolation of small RNA (< 200 b) and large RNA (> 200 b) in two separate fractions
Isolation of total RNA (small and large RNA in one fraction)
- Additional isolation of total protein fraction ready to use for SDS-PAGE and Western blot analysis
- Excellent RNA recovery and purity by chaotropic salt lysis without phenol/chloroform (patent pending)
- NucleoSpin® Filters for efficient sample homogenization
- rDNase for efficient on-column removal of genomic DNA
| Technology |
Silica-membrane technology |
| Format |
Mini spin columns |
| Sample material |
< 107 cultured cells, < 30 mg human / animal tissue
< 50 mg plant tissue, < 150 µl reaction mixture |
| Fragment |
size Small RNA: < 200 nt, large RNA: > 200 nt |
| Typical yield |
10 µg small RNA, 95 µg large RNA from 107 HeLa cells |
| Elution volume |
30–100 µl |
| Preparation time |
< 45 min/6 preps (small and large RNA)
< 35 min/6 preps (small RNA only) |
| Binding capacity |
200 µg |
Request a sample
Applications
- Parallel isolation of small and large RNA from human / animal tissue and cultured cells, from plant tissue, and in combination with phenol / chloroform (e.g., TRIzol®) lysis
- Purification of siRNA and large dsRNA from DICER reactions
- Typical downstream applications: real-time RT-PCR, Northern blotting, chip hybridization
Application data
 Very convenient RNA fractionation with highest selectivity
Very convenient RNA fractionation with highest selectivity
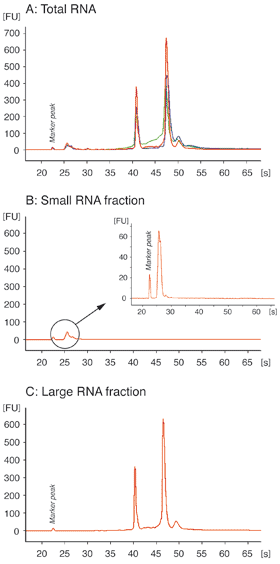
Total RNA was isolated from 107 HeLa cells using NucleoSpin® miRNA (red) and two competitor kits based on phenol/chloroform lysis and extraction (blue) or phenol/chloroform extraction (green). Equal amounts of total RNA fractions were analyzed on an Agilent Bioanalyzer (A).
NucleoSpin® miRNA allows isolation of small (B) and large (C) RNA in separate fractions in addition to the total RNA fraction (A).
NucleoSpin® miRNA allows selective isolation of small or large or total RNA fractions with very high recovery without the need of phenol/chloroform.
 Highly efficient removal of genomic DNA by on-column DNase digestion
Highly efficient removal of genomic DNA by on-column DNase digestion
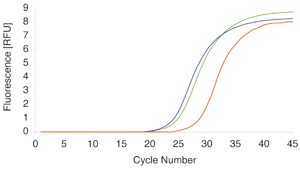
Total RNA from 107 HeLa cells was purified with NucleoSpin® miRNA (red) and two competitor kits Q (blue) and A (green). The RNA was assayed for residual traces of DNA by amplifying a 200 bp fragment of the ATPase 6 gene.
The ΔCts of 3.5 and 4.3 between NucleoSpin® miRNA and competitor A or Q, respectively, indicate a more than tenfold increase in DNA removal by the NucleoSpin® miRNA on-column DNase digestion compared to standard phenol/chloroform extractions.
 Direct linear correlation of input cells to RNA yield shown by quantitative RT-PCR
Direct linear correlation of input cells to RNA yield shown by quantitative RT-PCR
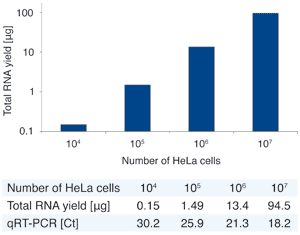
Total RNA was purified from 104, 105, 106, and 107 HeLa cells using NucleoSpin® miRNA. MiR-16 was amplified in a qRT-PCR reaction using a Roche LightCycler™.
The graph shows a perfect linear correlation of input cells and RNA yield. The data table additionally shows corresponding linear decrease of Ct values with increasing RNA input.
Excellent yields for all types of sample materials
Total RNA and fractionated RNA from maximum amounts of different sample materials were purified with NucleoSpin® miRNA according to the individual protocols. Note that RNA yields as well as the ratio of small to large RNA vary due to species, developmental stage, etc.
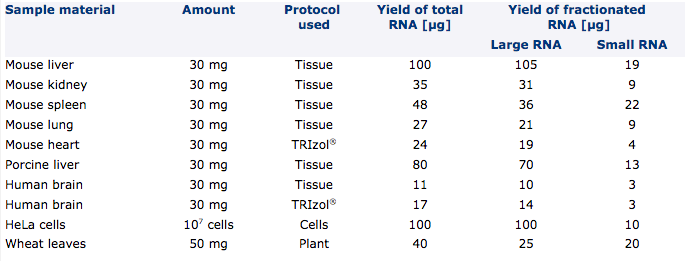
Cultured cells or soft tissue like liver, kidney, lung, etc. can easily be processed with the phenol-free standard procedures. For lipid tissue like brain or very hard-to-lyse, fibrous tissue like heart tissue it might be advantageous to use the protocol for RNA purification in combination with phenol/chloroform (e.g., TRIzol®) lysis to obtain optimal yields.
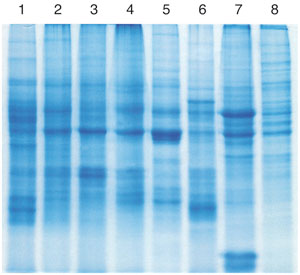 SDS-PAGE of precipitated protein fraction
SDS-PAGE of precipitated protein fraction
Total protein from various tissues (see table above)
was isolated during the NucleoSpin® miRNA procedure.
The protein precipitate was dissolved in Laemmli-like
protein solubilization buffer PSB and 40 µg were run on a 12% SDS polyacrylamide gel (100 V, 45 min).
1: Mouse liver
2: Mouse kidney
3: Mouse spleen
4: Mouse lung
5: Mouse heart
6: Porcine liver
7: Human brain
8: HeLa cells
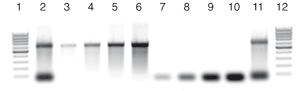 Purification of siRNA from a DICER reaction mixture
Purification of siRNA from a DICER reaction mixture
2.5, 5, 10, and 15 µg of a double stranded RNA template were digested using DICER enzyme (digestion efficiency was 60%).
The small siRNA product was purified from the reaction mixture by separating the uncut large dsRNA using the NucleoSpin® miRNA clean-up protocol for DICER reactions. The purified siRNA and large dsRNA fractions were analyzed on a 1% TAE agarose gel.
The yields of siRNA in lane 9 and large dsRNA in lane 5 correspond to the input amount (lanes 2 and 11) and show the high recovery of > 95%
1, 12: 100 bp DNA marker
2, 11: 25 µL (2.5 µg) unpurified DICER reaction mix
3–6: 25 µL purified large dsRNA fraction
(total elution volume: 100 µL)
7–10: 25 µL purified siRNA fraction
(total elution volume: 100 µL)
Please Note
The contents and the protocols of this kit have changed from July 2017! The kit will only contain blue NucleoSpin RNA Columns. The protocols will only show the collection of total RNA in one fraction. There are support protocols available for the fractionation of small and large RNA. Contact us if you have any questions.
|
Request a sample
 Very convenient RNA fractionation with highest selectivity
Very convenient RNA fractionation with highest selectivity Highly efficient removal of genomic DNA by on-column DNase digestion
Highly efficient removal of genomic DNA by on-column DNase digestion Direct linear correlation of input cells to RNA yield shown by quantitative RT-PCR
Direct linear correlation of input cells to RNA yield shown by quantitative RT-PCR 
 SDS-PAGE of precipitated protein fraction
SDS-PAGE of precipitated protein fraction Purification of siRNA from a DICER reaction mixture
Purification of siRNA from a DICER reaction mixture