Find your product
Advanced searchLogin
Webshop under construction
Due to technical maintenance the webshop is closed until January 3, 4. We wish you a successful 2024!
Welcome
Sign outPersonal information

NEBNext Direct for target enrichment
NEBNext Direct® employs a unique technology that enables highly specific target enrichment of genomic regions of interest. The result of a partnership between New England Biolabs® (NEB®) and Directed Genomics™, this innovative approach to target enrichment balances the speed and precision of multiplexed PCR-based approaches with the content scalability typical of hybridization-based methods. This flexibility allows a single workflow for assays ranging from single gene tests to comprehensive panels including several hundred genes. Regardless of sample type or assay content, NEBNext Direct allows you to enrich your targets with precision. Read what other people are saying.
Advantages
- Generate a higher percentage of your sequencing reads aligning to your targets
- Obtain uniform sequencing of ALL targets, regardless of GC content
- Save time with a streamlined workflow that couples enrichment and library preparation
- Generate high quality libraries with limited input amounts and degraded DNA samples, including FFPE and ctDNA
- Distinguish molecular duplicates, reducing false positive variants and improving sensitivity
- On-bead sample preparation is ideal for automated workflows
Request More info or a free sample
NEW: NEBNext Direct Custom Ready Panels
Employing the unique NEBNext Direct hybridization-based enrichment method, NEBNext Direct Custom Ready Panels allow rapid customization of targeted gene panels for Illumina sequencing. Select from a list of genes for which baits have been carefully designed and optimized to give complete coverage of the full coding regions. High quality panels can be designed by you and rapidly delivered, from any combination of genes. NEBNext Direct Custom Ready Panels provide the content you want with the performance you need.
Advantages
- Choose from a single gene to hundreds of genes
- Experience unmatched specificity and coverage uniformity
- Eliminate synthesis and optimization steps for faster turnaround
- Improve sensitivity with our Unique Molecule Index (UMI)
- Generate results in one day with our automation-friendly workflow
More information about the NEBNext Custom panels
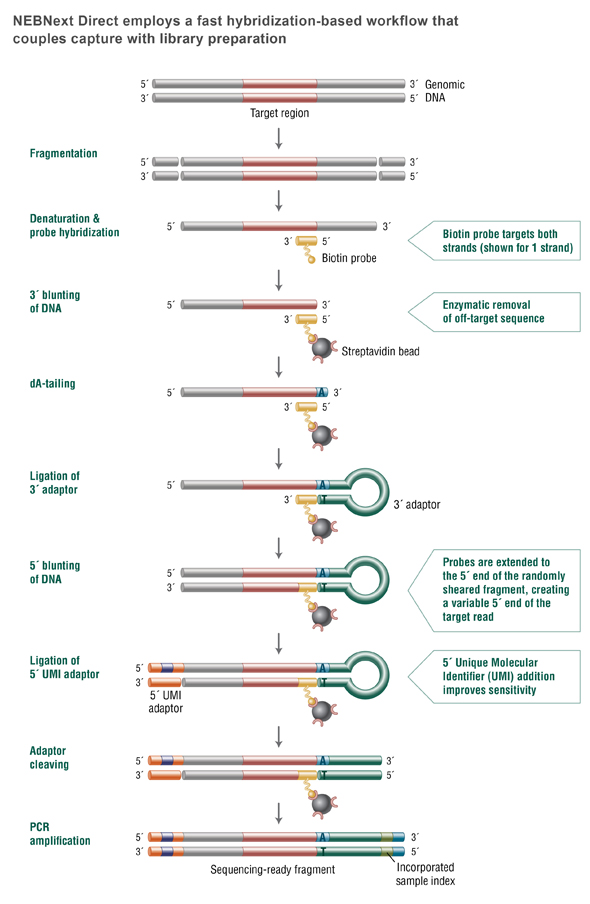
NEBNext Direct Workflow
View our tutorial for help visualizing NEBNext Directs's unique and enabling workflow:
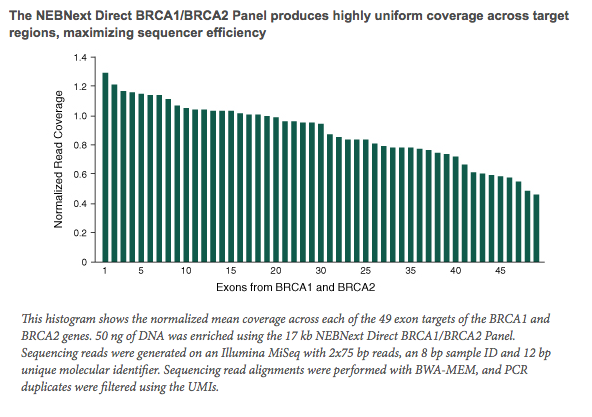
- The NEBNext Direct BRCA1/BRCA2 Panel
- Try NEBNext Direct® for yourself. Request a NEBNext Direct sample.
- For more information, please fill in the form
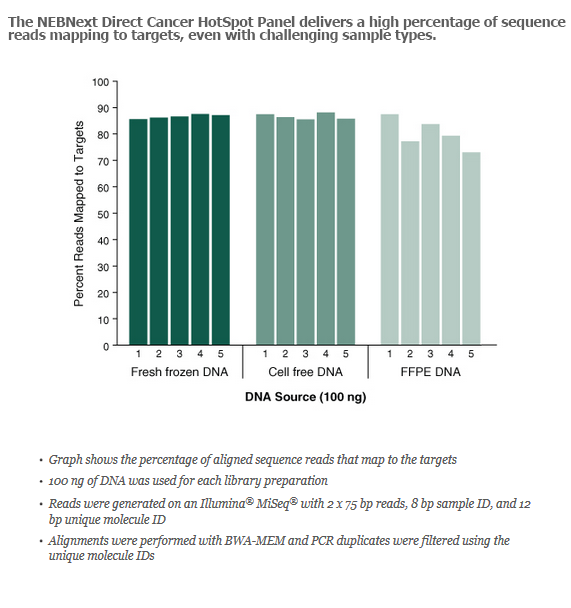
- Learn more about the NEBNext Direct Cancer HotSpot Panel and view performance data.
- View questions and answers
What People are Saying
"NEBNext Direct enrichment technology is by far the fastest and most automation friendly protocol available today. I can have samples on the sequencer in 6 hours starting from genomic DNA. The technology produces very high on target percentages (>90%) for even very small panels, and in combination with molecular barcoding produces low duplication rates. From an optimization perspective, NEBNext Direct enrichment allows me to assign individual captured fragments to a probe unambiguously, thus giving the opportunity for optimizing the coverage distribution of any target."
– Eric C. Olivares, Founder, SEQanswers.com
"Using the NEBNext Direct kit, we were able to detect all known single nucleotide variants and indels in DNA extracted from fresh frozen or FFPE tissue derived from glioma biopsies. We could also clearly see amplification in genes like EGFR or PDGFR. The workflow is really easy and fast and can be rapidly implemented in a lab."
– Yannick Marie, Sequencing Core Facility Manager, Brain and Spine Institute (ICM)
"The kit and its technology are easy to use and easy to automate, allowing us to get up and running quickly. The protocol itself is fast and efficient to obtain deep coverage of targets, giving homogeneous results for FFPE and frozen tumors, therefore opening doors for customized panels."
– Francis Rousseau, Ph.D. Director of Genomics for IntegraGen SA